- Details
Projects
The group is at the intersection of life and computational sciences, with a main focus on biological processes in motion such as cell migration and disease dynamics.
We make extensive usage of signal processing and bioimage analysis to study complex cellular dynamics, with a main focus on leukocyte motility and interaction. We also translate these techniques to different domains such as medical imaging, digital screening, and longitudinal patient monitoring through wearables.
The group is distributed between the TKI of University of Bern (with a wet-lab part), the Euler institute of USI Lugano (with a dry-lab part), and is affiliated to the International Center for Advanced Computing in Medicine (ICAM) of University of Pavia.
We are happy to host interns MSc theses and visiting PhD students to pursue an interdisciplinary training.
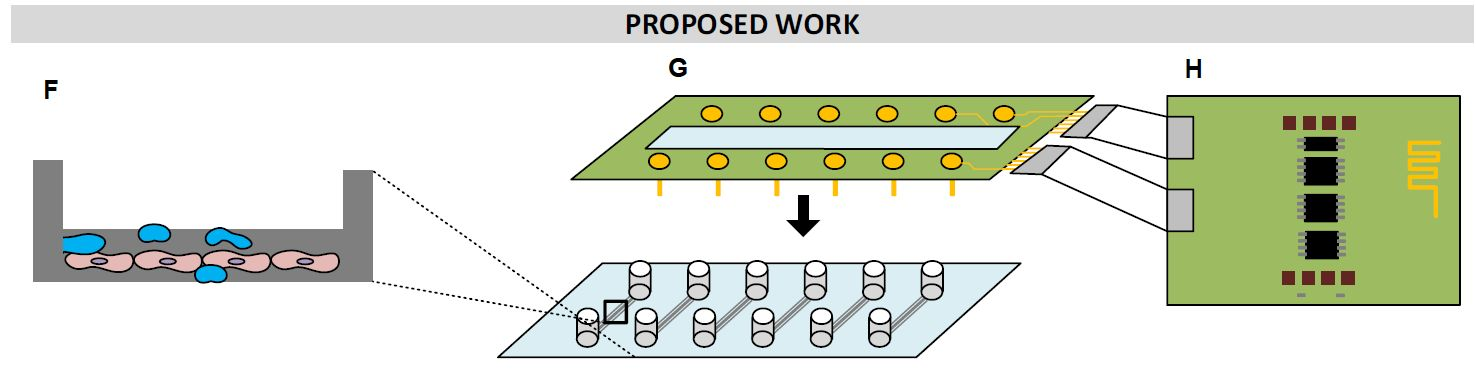
EXTRAVOLT: Modulating extravasation of immune cells across the brain-blood-barrier via electric stimulation
Electric signals regulate a variety of biological processes in our body. However, their interaction with the immune system remains largely unknown. Supported through a SNSF SPARK grant, we are pioneering the field of “electro-immunology”, exploring if and how inflammatory immune cells respond to electric stimulation with the aim of inhibiting their infiltration in the central nervous system.
Leveraging the expertise available at the TKI, including in-house microfluidic models of the blood-brain-barrier (BBB), we are specifically testing the effect of electric stimulation to modulate the extravasation of inflammatory T cells through the BBB. If successful, this project could set the basis for a new therapeutic strategy for neuroinflammation based on electricity, which has the advantage to be delivered locally and on demand.
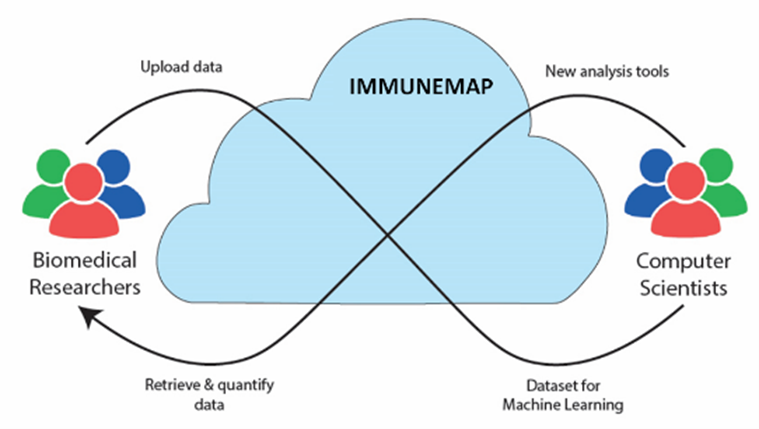
IMMUNEMAP: an open atlas of immune cell motility
This project, co-directed by the group together with Prof. Gonzalez (IRB Bellinzona) and Prof. Krause (KAUST), aims at disseminating Open Research Data practices in bioimaging, providing an open platform to share and promote the reusage of intravital microscopy videos capturing immune system dynamics. This is essential to bridge the gap between researchers in immunology and computer vision, enabling exchange of data, analysis algorithms and reciprocal needs.
Thanks to an international network of 20 laboratories (including TKI), we are providing 1M+ single cell annotations, 50k+ cell trajectories and 400+ intravital microscopy videos acquired in different anatomical regions and experimental conditions.
Leveraging immunemap and unsupervised machine learning algorithms, we uncovered specific motility patterns displayed by immune cells in different experimental conditions.
Link: www.immunemap.org
Cell Behavior Video Classification Challenge (CBVCC)


The primary goal of the CBVCC is to create models that can accurately classify videos based on the movement patterns of cells. Specifically, the models should be able of:
- Identifying videos where cells exhibit sudden changes in their direction of movement.
- Distinguishing these from videos where cells show consistent, linear movement, stationary cells as well as from videos containing only background.
Rules, registration and details: https://immunemap.org/index.php/challenges-menu/cbvcc
Leaderboard: https://www.dp-lab.info/cbvcc
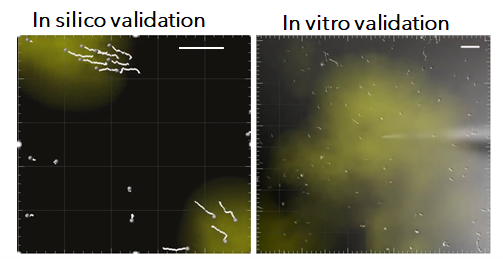
Visualizing invisible chemokine gradients
Chemokine gradients are key driver of interstitial cell migration. However, they are difficult to visualize and quantify in living tissue due to extremely low chemokine concentrations and their transient nature. Supported by a FIR grant, this project aims at estimating chemokine distribution maps from the movement of cells, potentially revealing chemokine sources and their effect on leukocyte motility.
By combining time-lapse microscopy with simulations of chemokine sources and cell motility developed in collaboration with Prof. Wortel (Radboud University – NL) we trained a deep learning method to estimate chemokine maps from cell tracks. Moreover, we are starting a collaboration with Prof. Cui (Harvard Medical School) to validate these findings through spatial transcriptomics. Preliminary results of this project were presented at the European Conference on Cell Migration, Leuven 2023 received the award “outstanding contribution to immunological research from EFIS/EJI”